Dose response
Scientific Background¶
To select a practical number of compounds that exhibited high activity in the primary screen for follow-up assays, a cutoff value of the primary %Inhibition was applied. The cutoff value was calculated as the sum of the average percent inhibition of all compounds tested and three times their standard deviation. In this case, 48 unique compounds (1.9% of all compounds screened) were selected as hits.
As before, the number of unique compounds selected for follow-up screening represents a relatively small sample size and therefore were immediately progressed to individual dose response analysis (ED50 determination).
Compounds were "hit-picked" at 10 mM concentration in DMSO and further serially diluted ten times at three-fold dilutions for a total of 10 points per compound. Data was normalized as in the primary assay and curves were plotted and fitted to a four-parameter equation describing a sigmoidal concentration-response curve (for example, the Hill equation). The reported ED50 values are generated from fitted curves by solving for the x-intercept at the 50% activity level of the Y-intercept. Compounds with ED50 values greater than 0.01 micromolar are considered inactive; compounds with ED50 of equal to or less than 0.01 micromolar are considered active.
Third-Party Software¶
This tutorial was developed using TIBCO Spotfire 11.4. Visualization templates may need adjustments if using older versions of that software.
Dataset¶
Prerequisite: Control Layout Dose Response must exist.
Six serial dilution plates were used. Minimum and maximum controls were placed in columns 1 and 12, respectively.
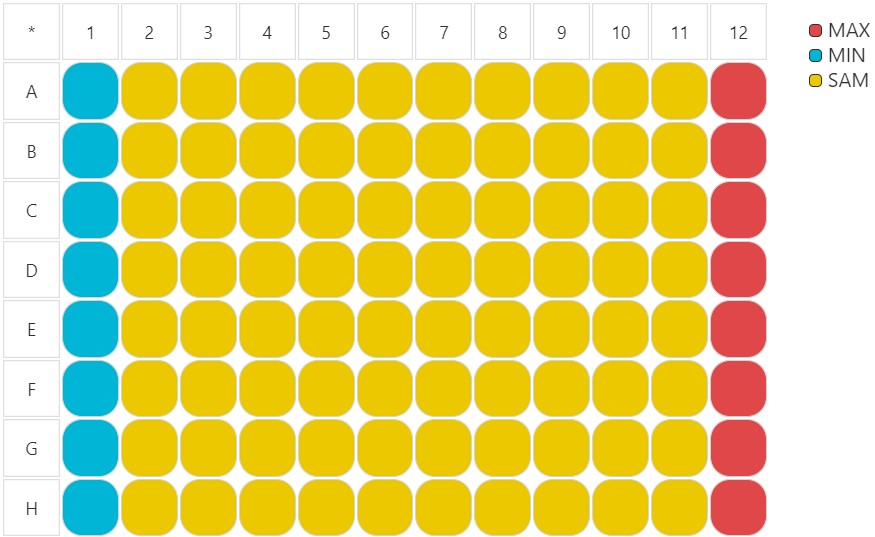

Raw data was generated by an AnalystGT Reader from Molecular Devices.
Files to use:
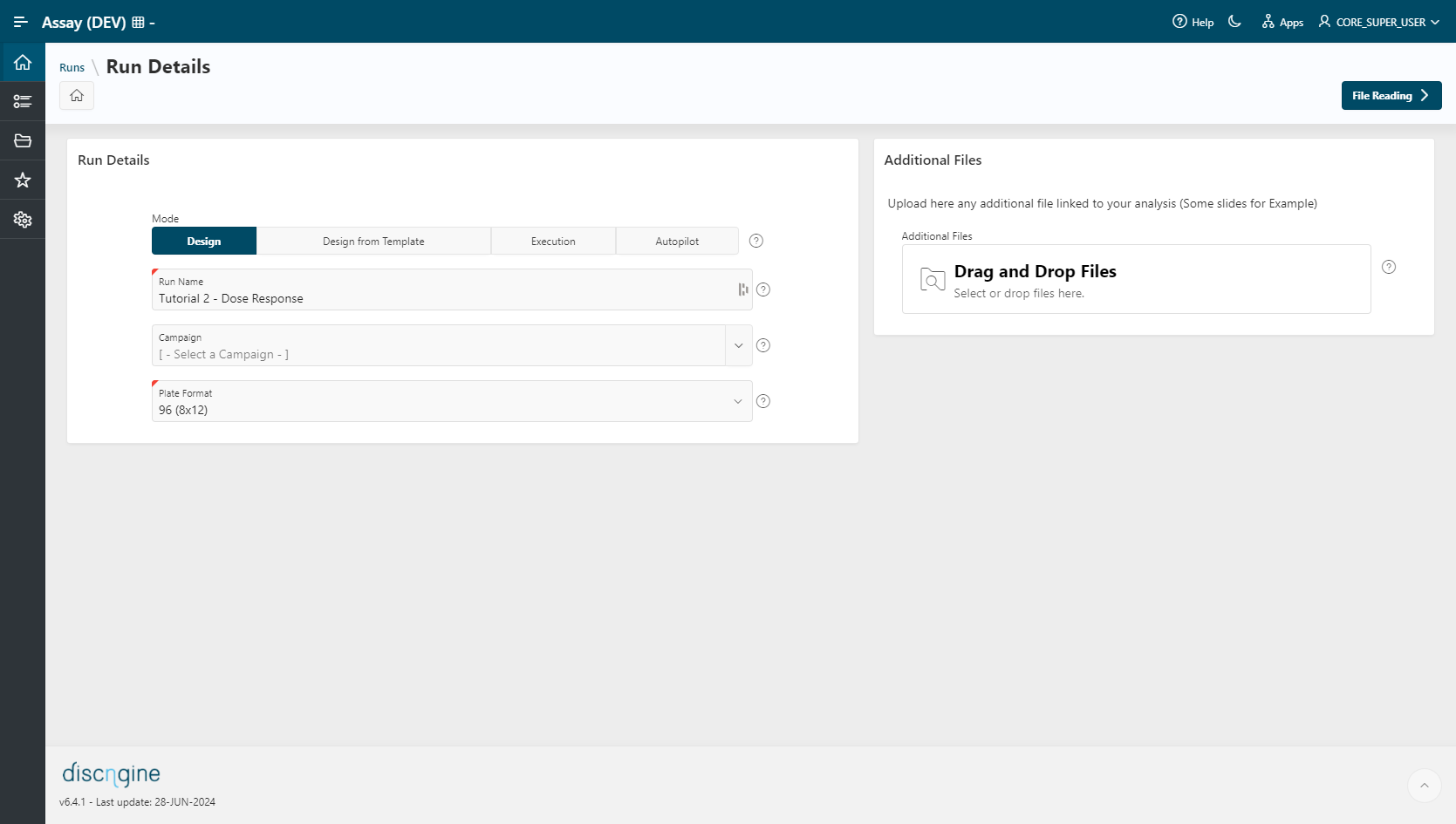
Run Creation¶
- To create a new run, click the
New Runbutton. - Select Mode as
Design Mode. - Enter a
Run Name. - Specify the plate format to use:
96 (8x12). - Go to the File Reading step.
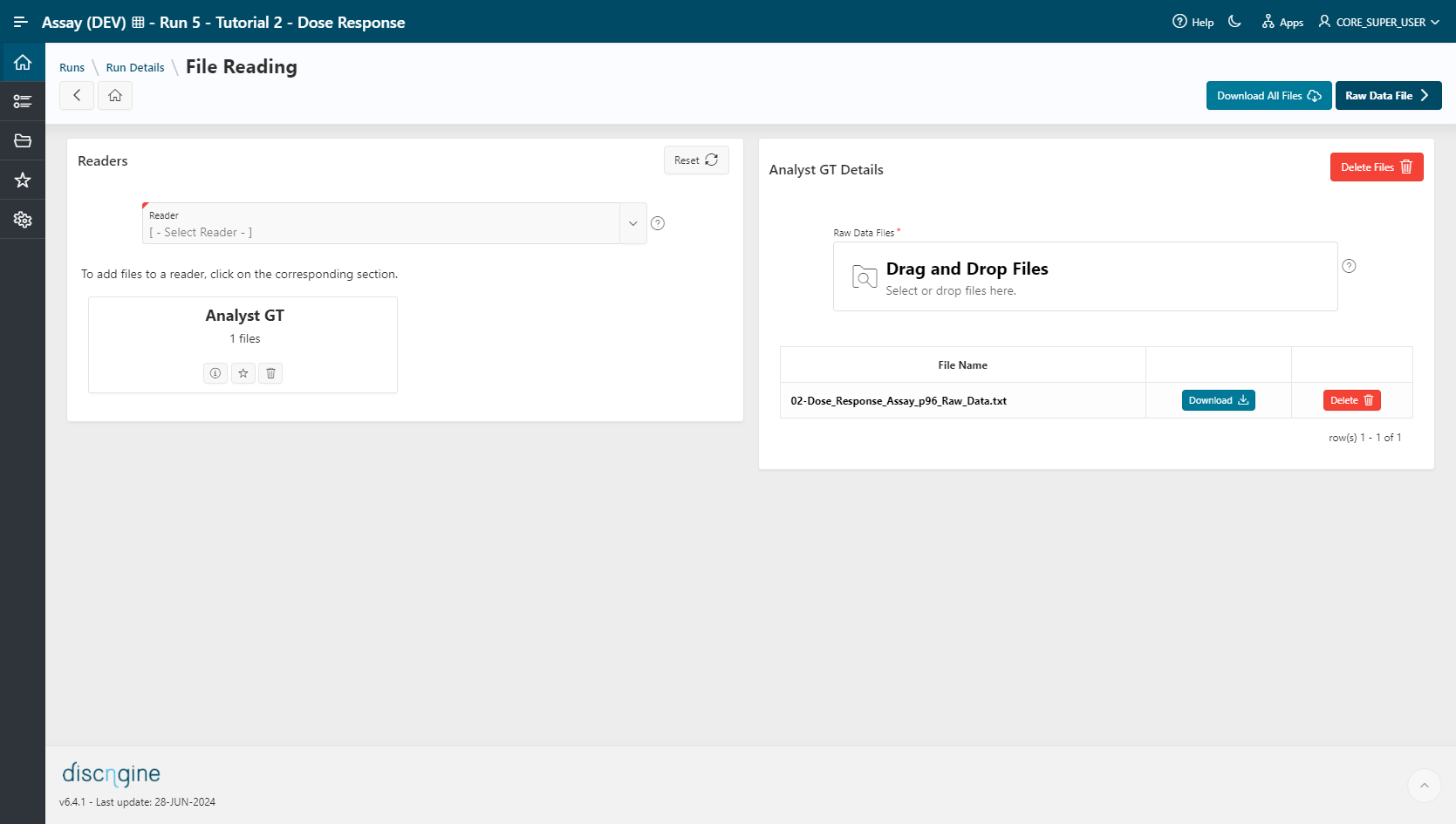
File Reading¶
- Select
Analyst GT (TAB)as a reader. - Select the file
02-Dose_Response_Assay_p96_Raw_Datain the upload section. The file is automatically uploaded upon selection and listed in the table below. - Go to the Raw Data File step.
Raw Data File¶
- The Number of Measurements per Plate (
1) and the READOUT Alias (READOUT1) are already set. - Go to the Plate Mapping step.
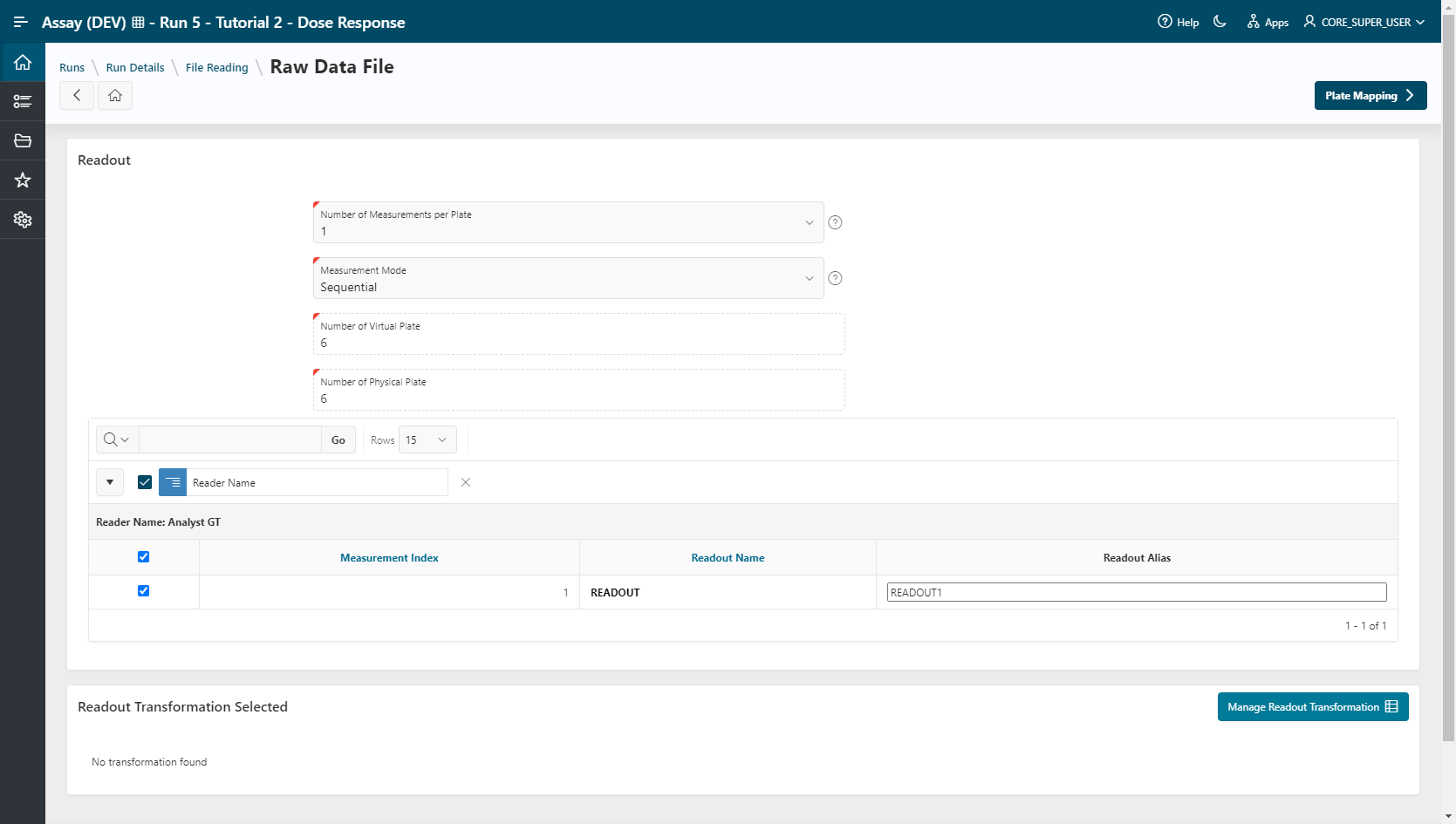
Plate Mapping¶
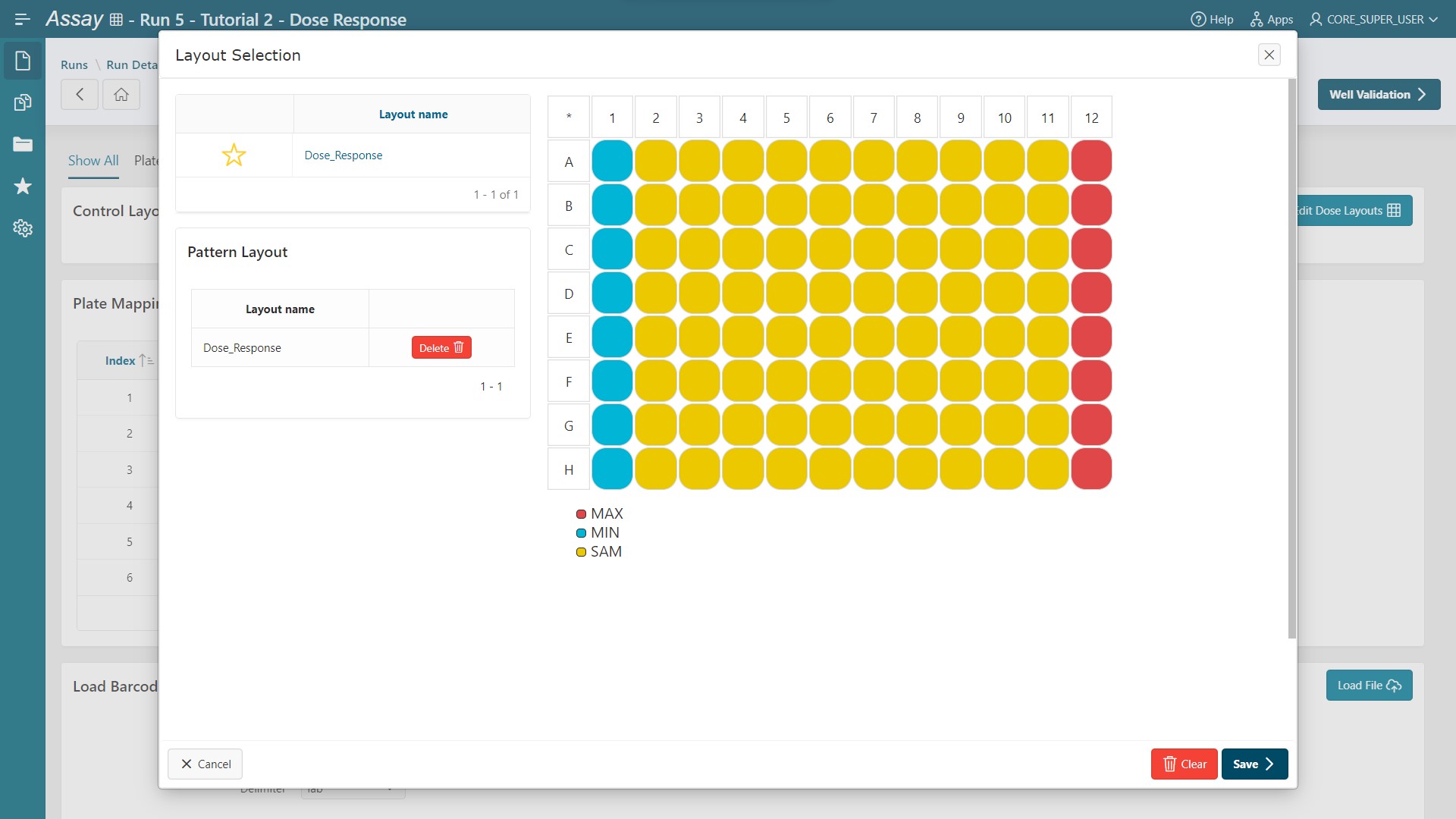
- Click
Edit Control Layouts. - Select the Control Layout named
Dose_Response(Minimum control in column 1 and maximum control in column 12). - Click
Save.
Info
Barcode information is not needed in this tutorial. This section allows users to set plate barcodes using a file with two columns (Plate_Index / Barcode).
- All your plates have the
Dose_Responselayout.
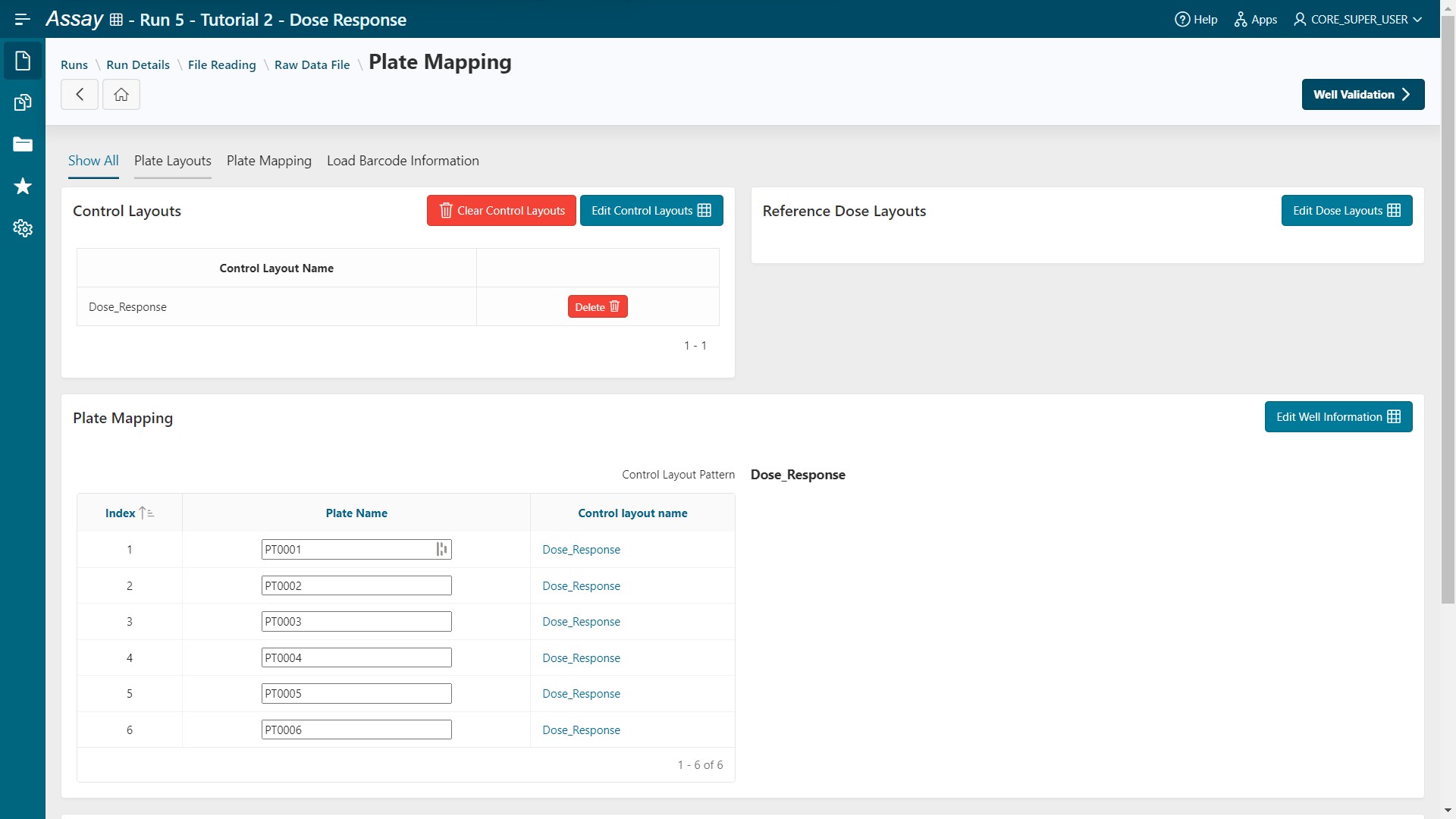
- Go to the Well Validation step.
Well Validation¶
Select your visualization tool:
Info
If you are using Spotfire Analyst Client, or if only one visualization tool is installed in your environment, this step will be skipped.
- TIBCO Spotfire Webplayer (Template: ValidationDefaultTemplate)
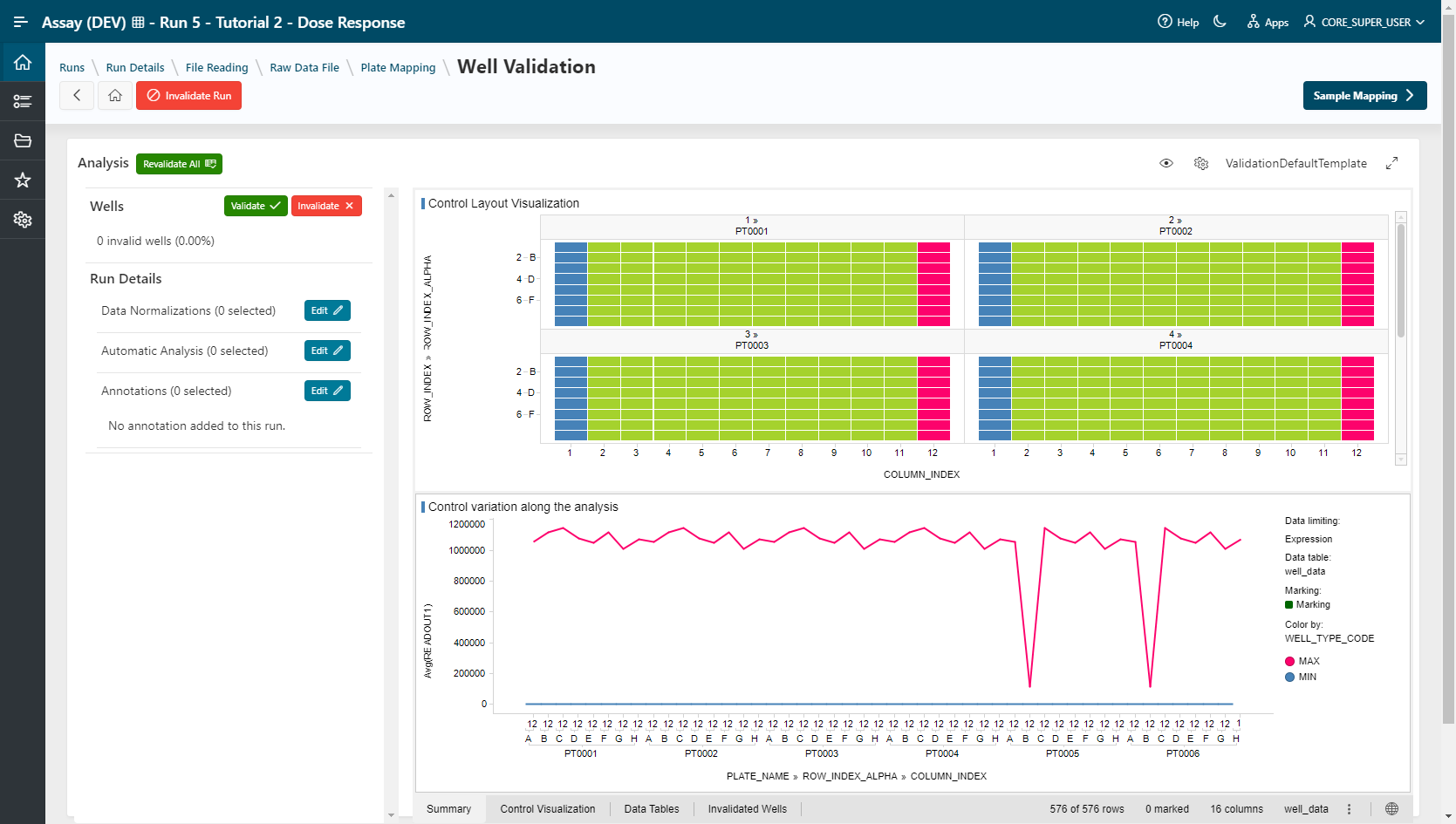
- Pipeline Pilot (Template: Well Validation)
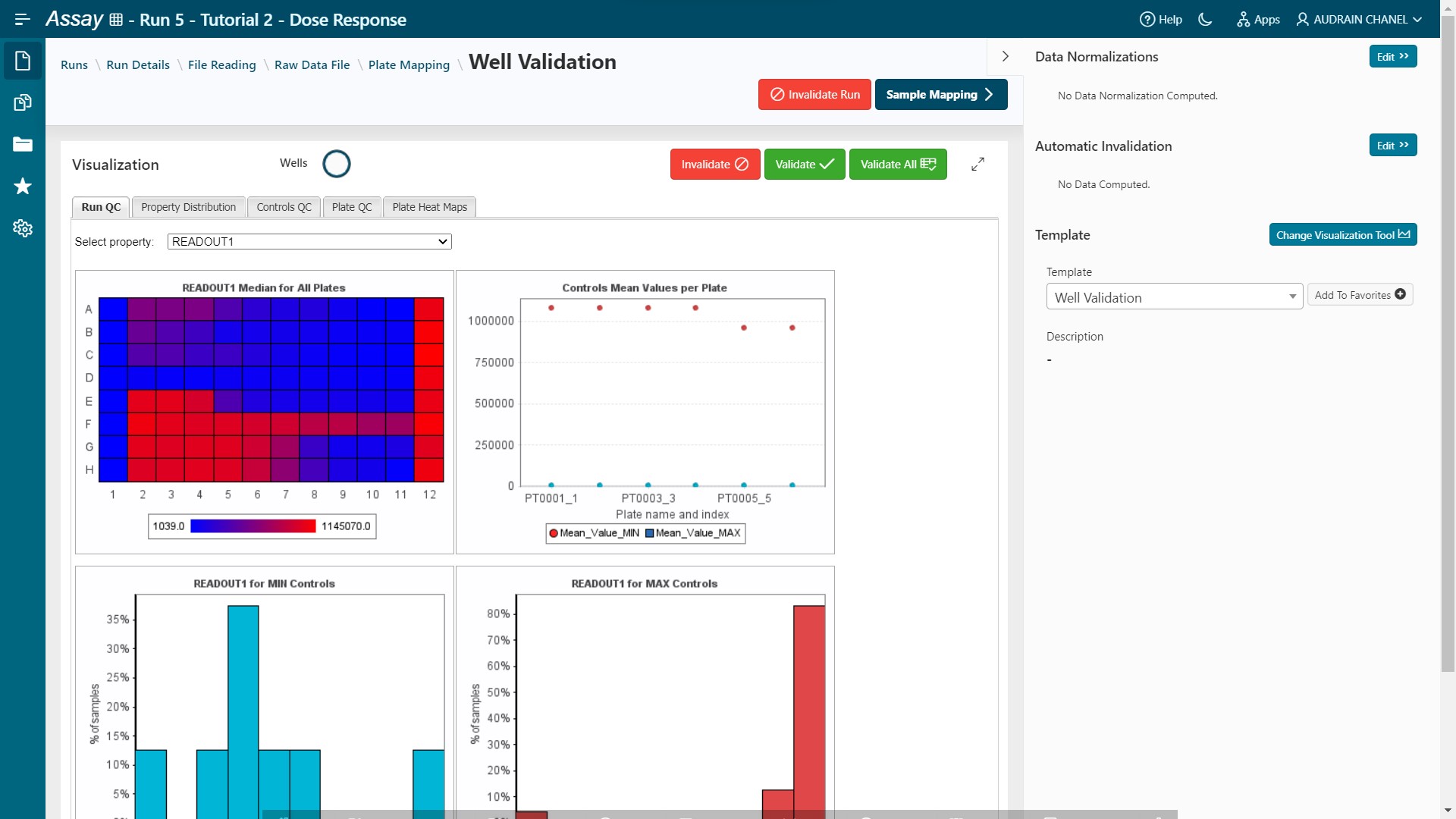
Results are loaded.
Add Data Normalizations:
- In the left panel, next to the
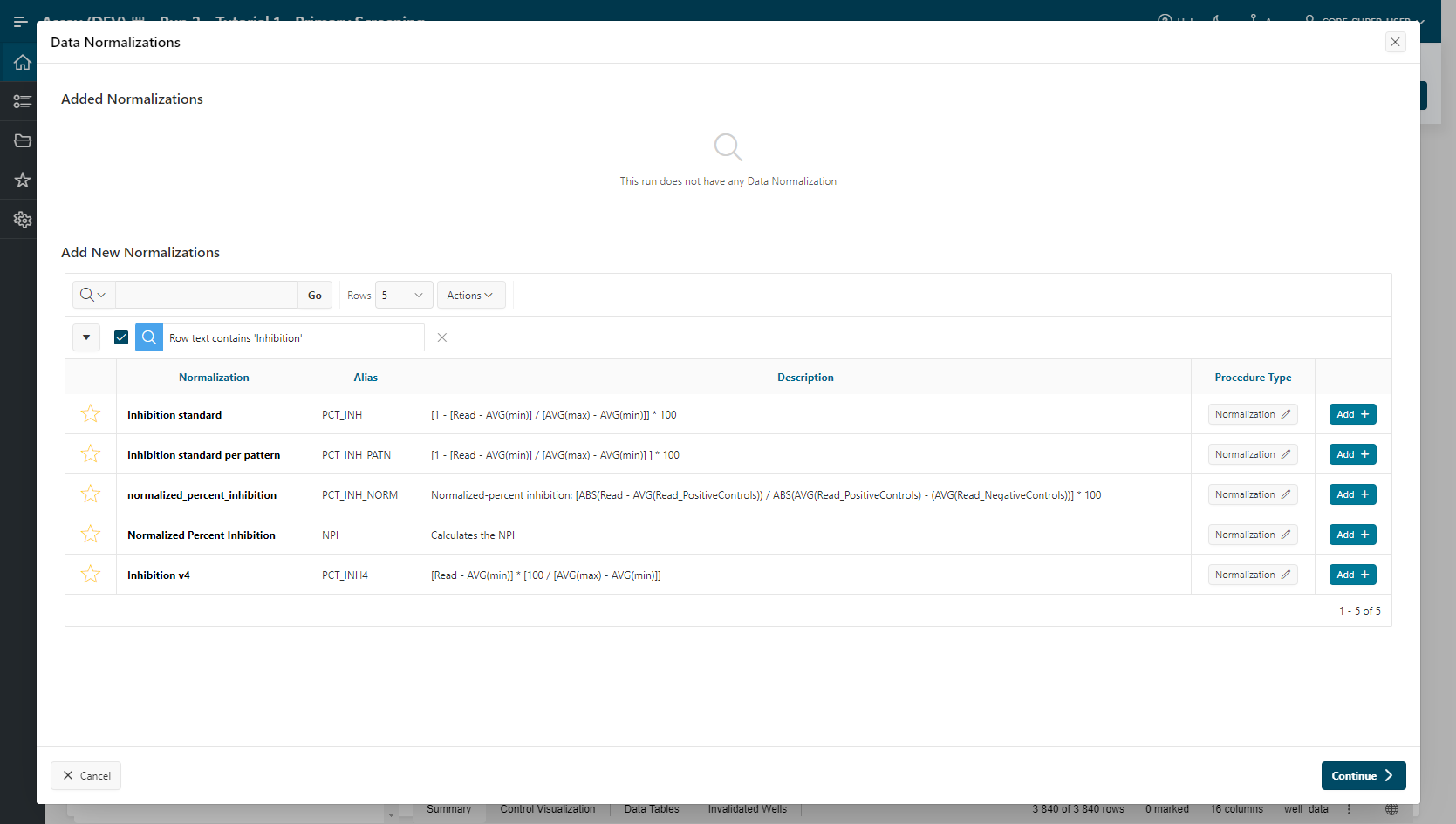
Data Normalizationsummary, clickEditto open the Data Normalizations modal. - Look for the
Inhibition Standardprocedure and clickAdd:- Value
READOUT1,MIN, andMAXare selected. - Click
Add.
- Value
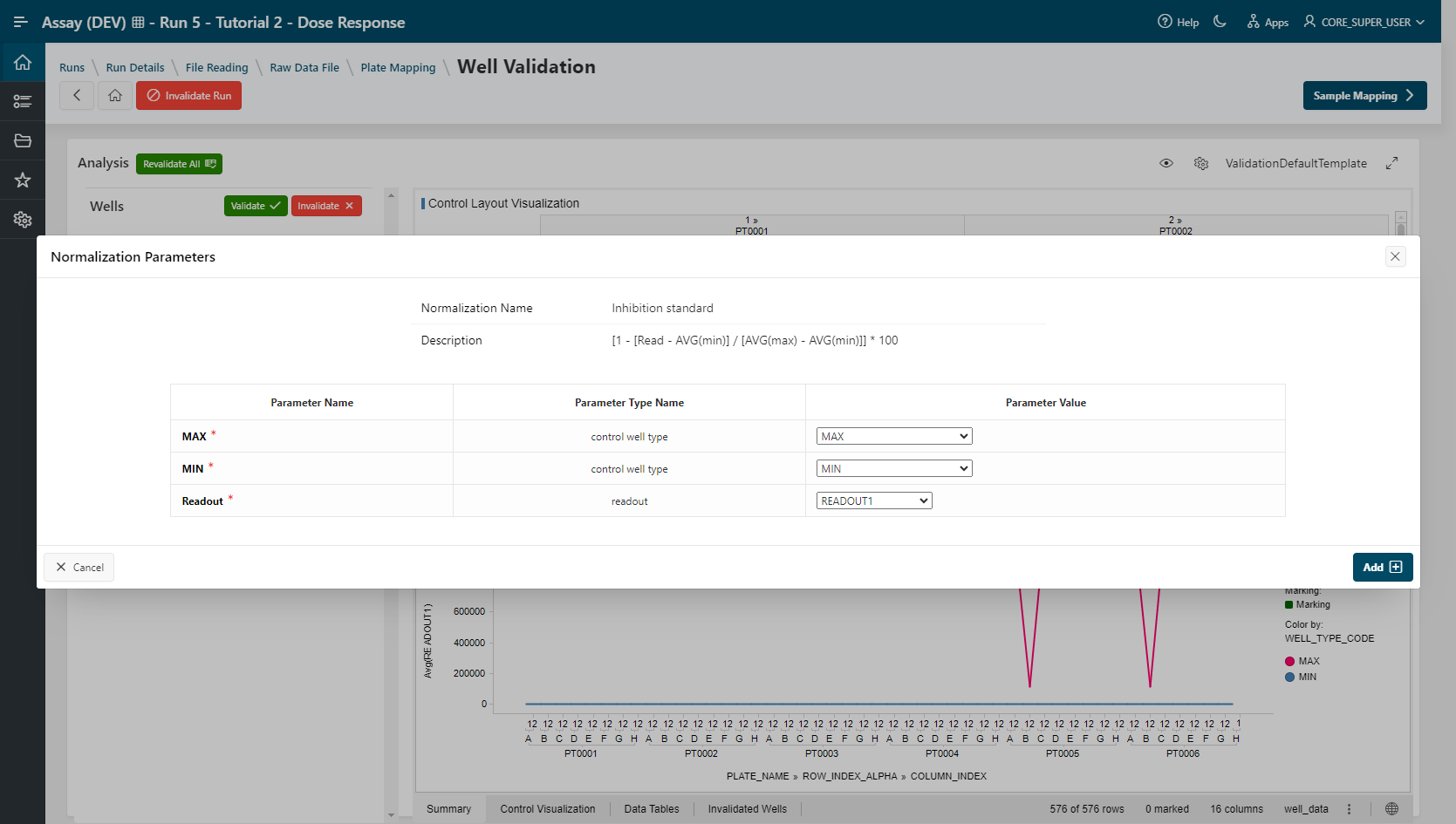
The Inhibition Standard normalization is listed in the Data Normalization Summary section.
- Repeat this action for Plate Z'-factor normalization:
- Values
READOUT1,MIN, andMAXare selected. - Click
Add.
- Values
The Plate Z-factor normalization is listed in the Data Normalization Summary section.
- Repeat this action for Z-Score per Type (Plate) normalization.
- Value
READOUT1is selected. - Click
Add.
- Value
The Z-Score per Type (Plate) normalization is listed in the Data Normalization Summary section.
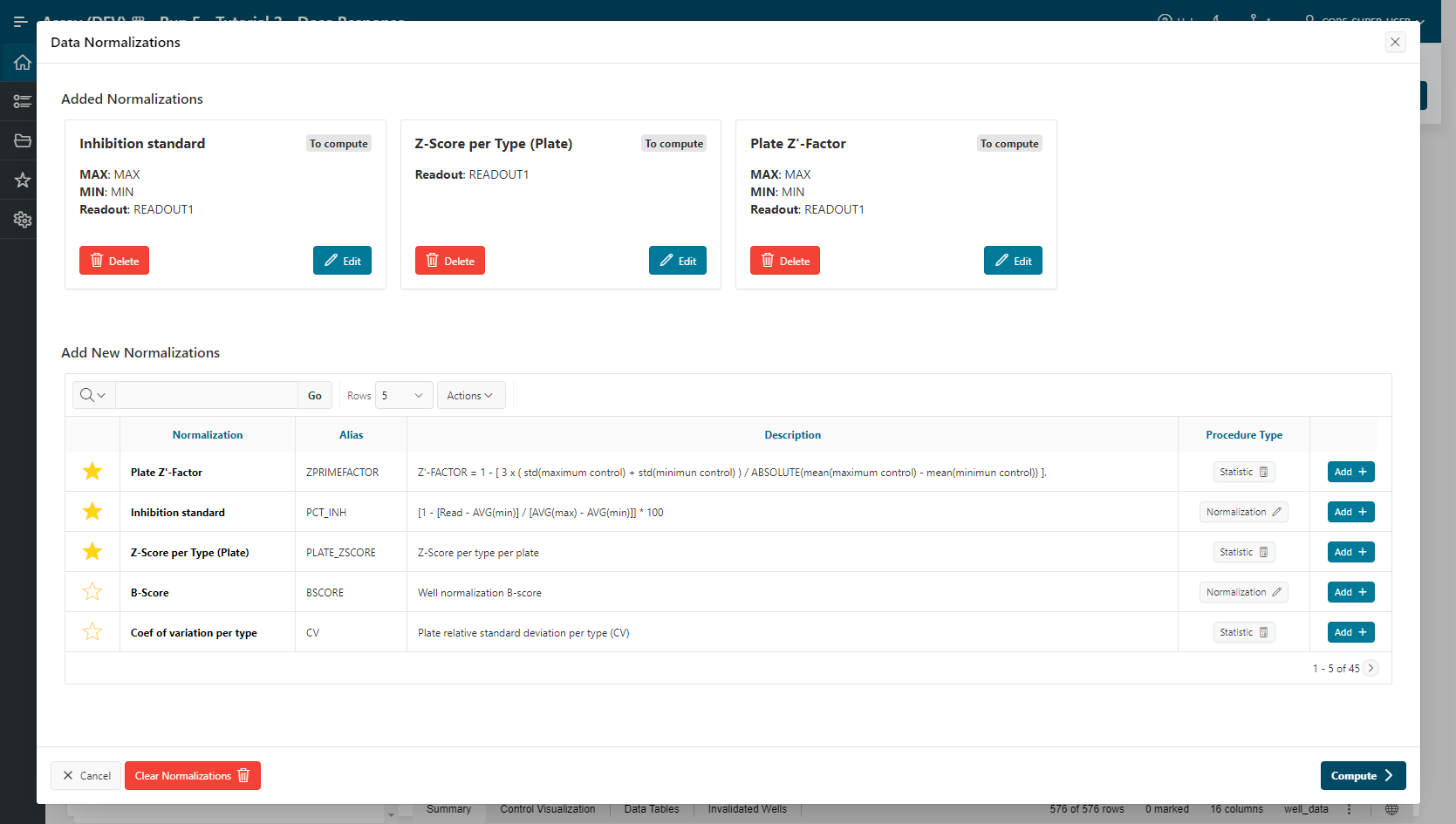
- Click on
Compute. You are automatically redirected to the Well Validation page.
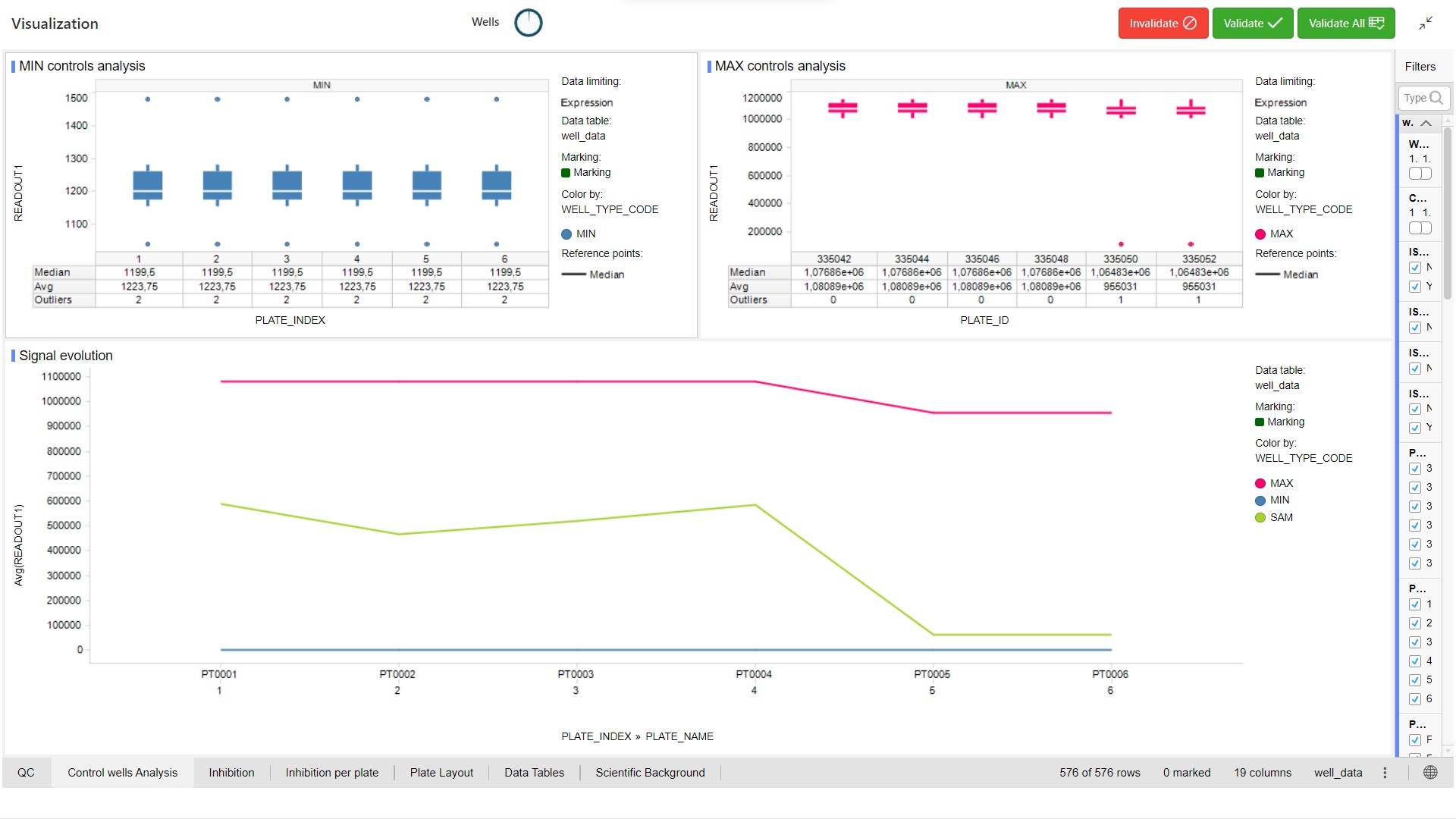
All normalizations are processed. Select another visualization template by clicking on the gears button above the current visualization.
- Apply the Dose_Response_Template (Spotfire)
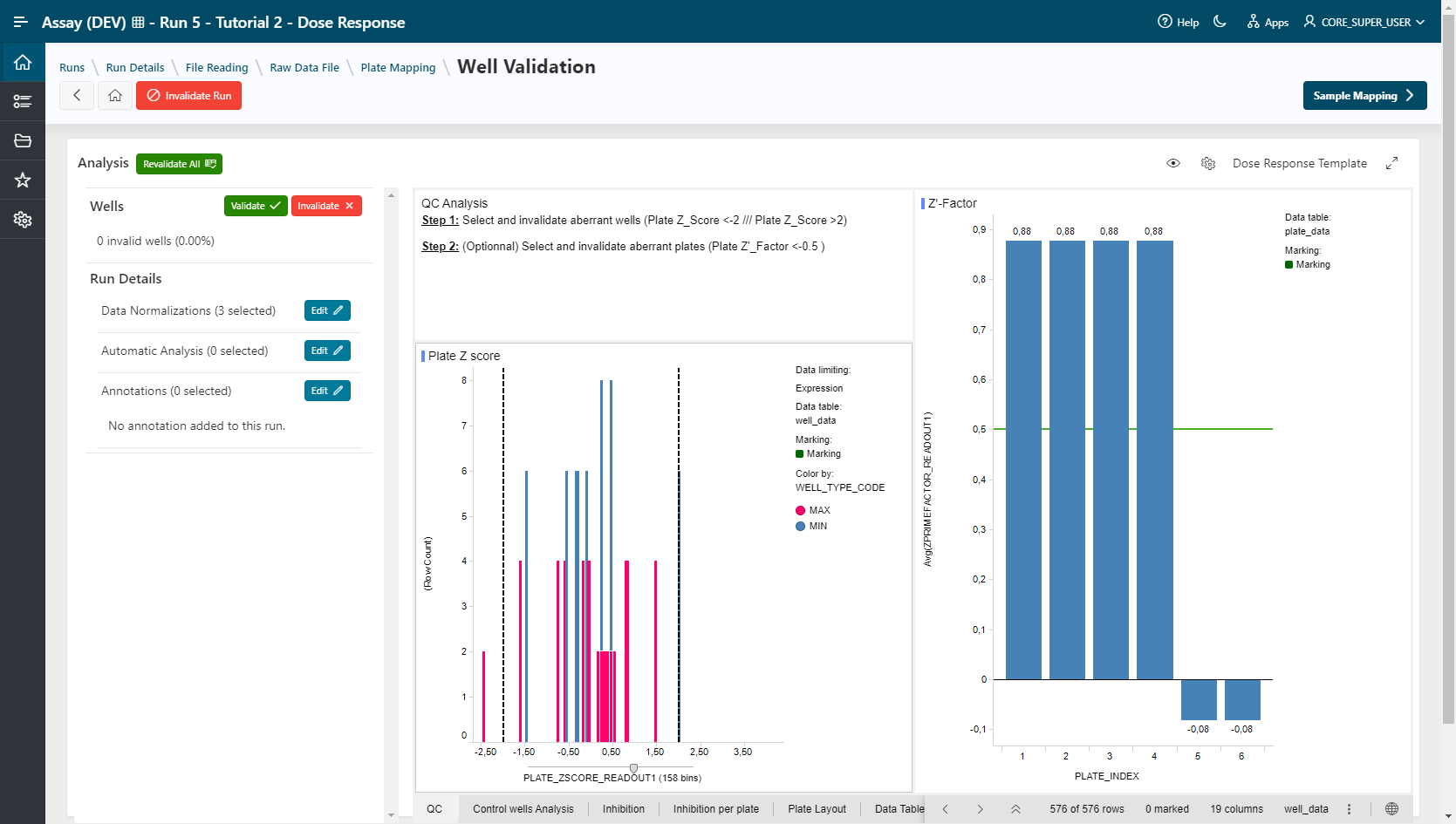
- Apply the Dose Response Template (Pipeline Pilot)
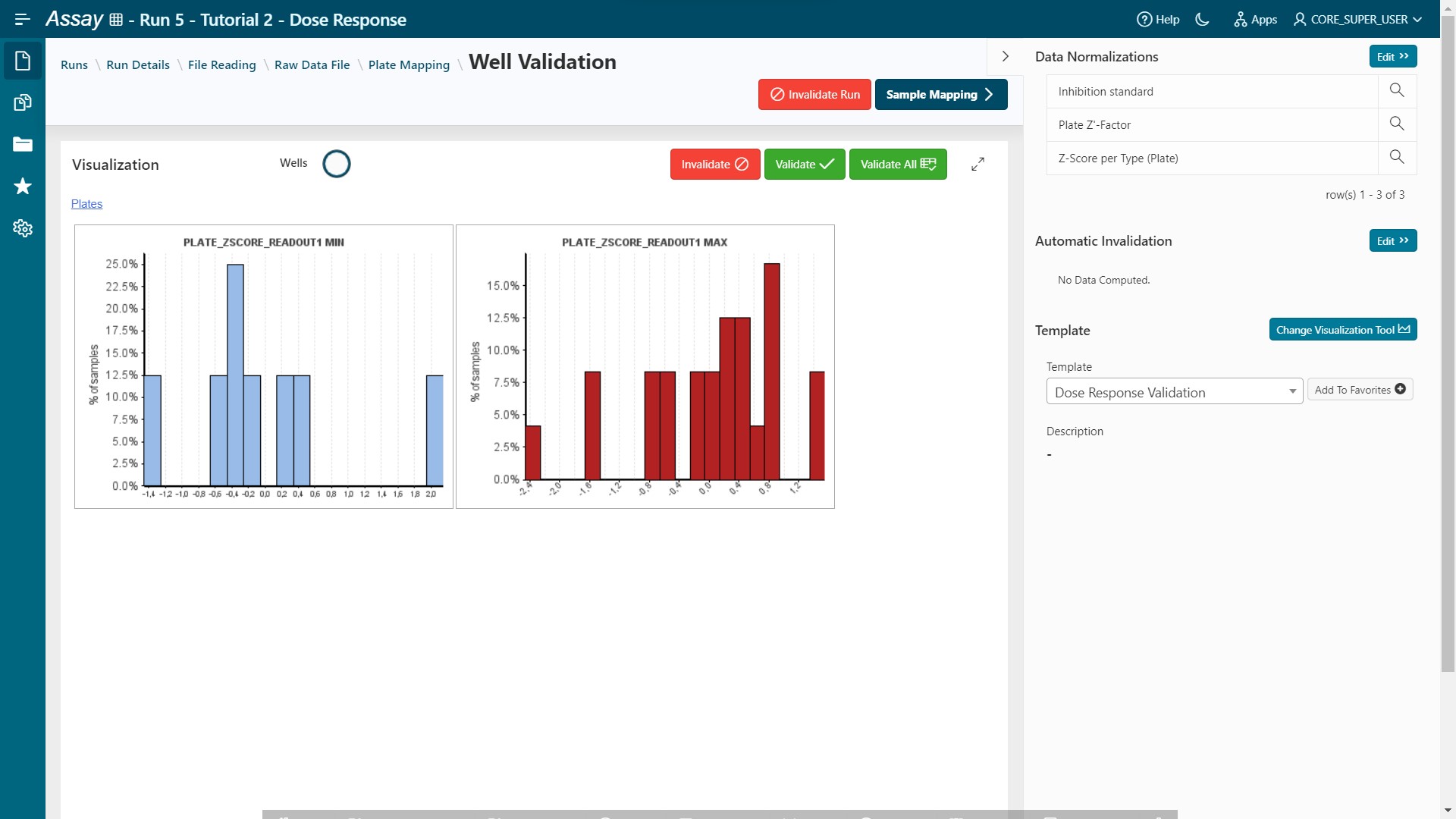
- Or create a new one and save it (see developer guide to register a new Pipeline Pilot Protocol in Assay).
For QC Validation, select aberrant wells in the visualization and click on the  button. Multiple selection is allowed by holding the Ctrl key.
button. Multiple selection is allowed by holding the Ctrl key.
After each invalidation, the normalizations will be relaunched in order to recalculate based on your selection.
Example
Plate_ZSCORE_READOUT1 < -2 or Plate_ZSCORE_READOUT1 > 2. (8 wells are invalidated) and all normalizations are recomputed.
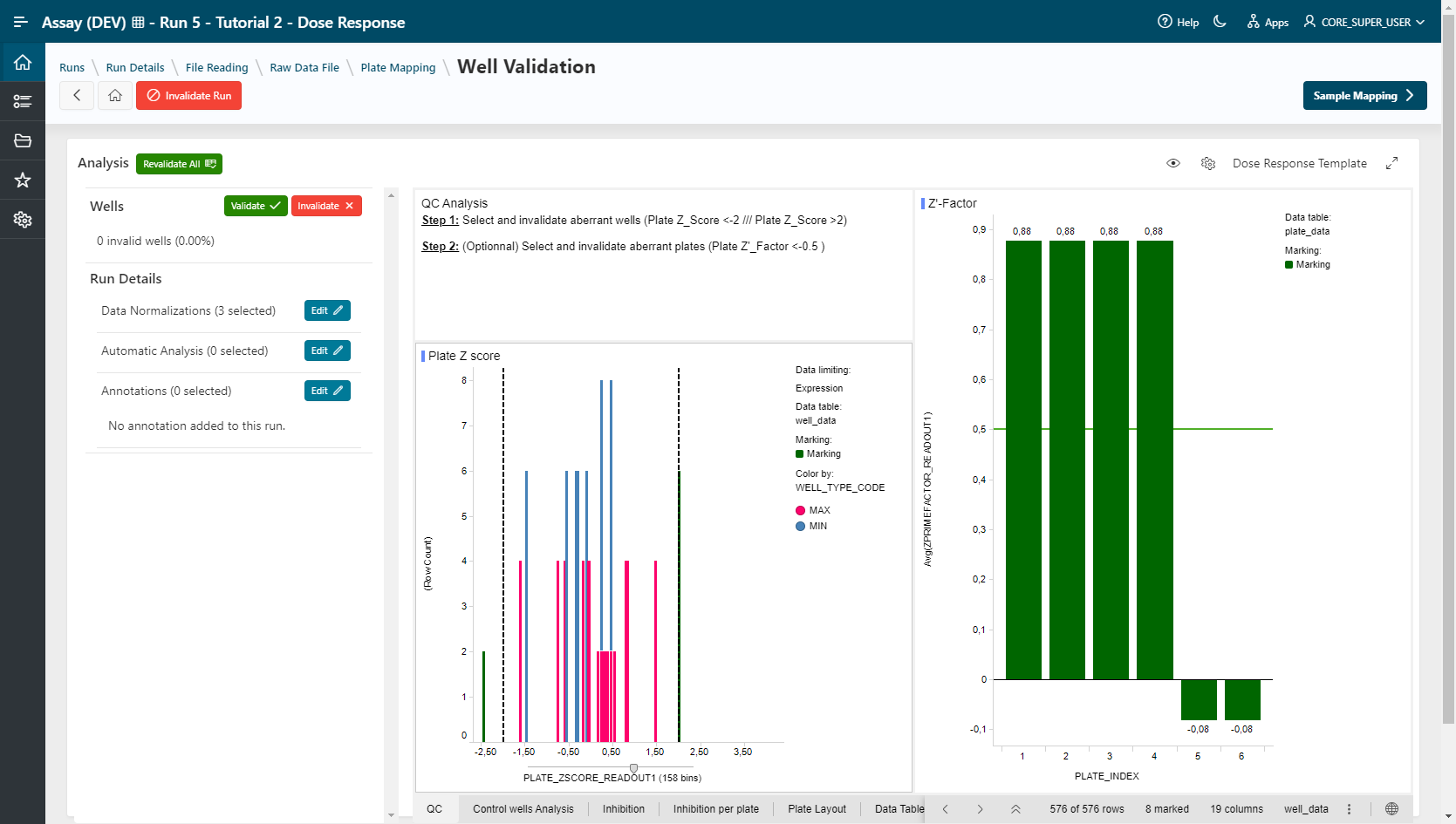
Tip
The space occupied by the visualization can be optimized to your liking by clicking on the  icon and the
icon and the 
- Go to the Sample Mapping step
Sample Mapping¶
- Click on
Edit Sample Informationbutton. - Select filling mode
Fileand tickSampleandDosefor content. - Load the file
02-Sample_Mapping_Dose_Response_Assay_p96. - Click on
Applybutton. Samples and doses are filled in.
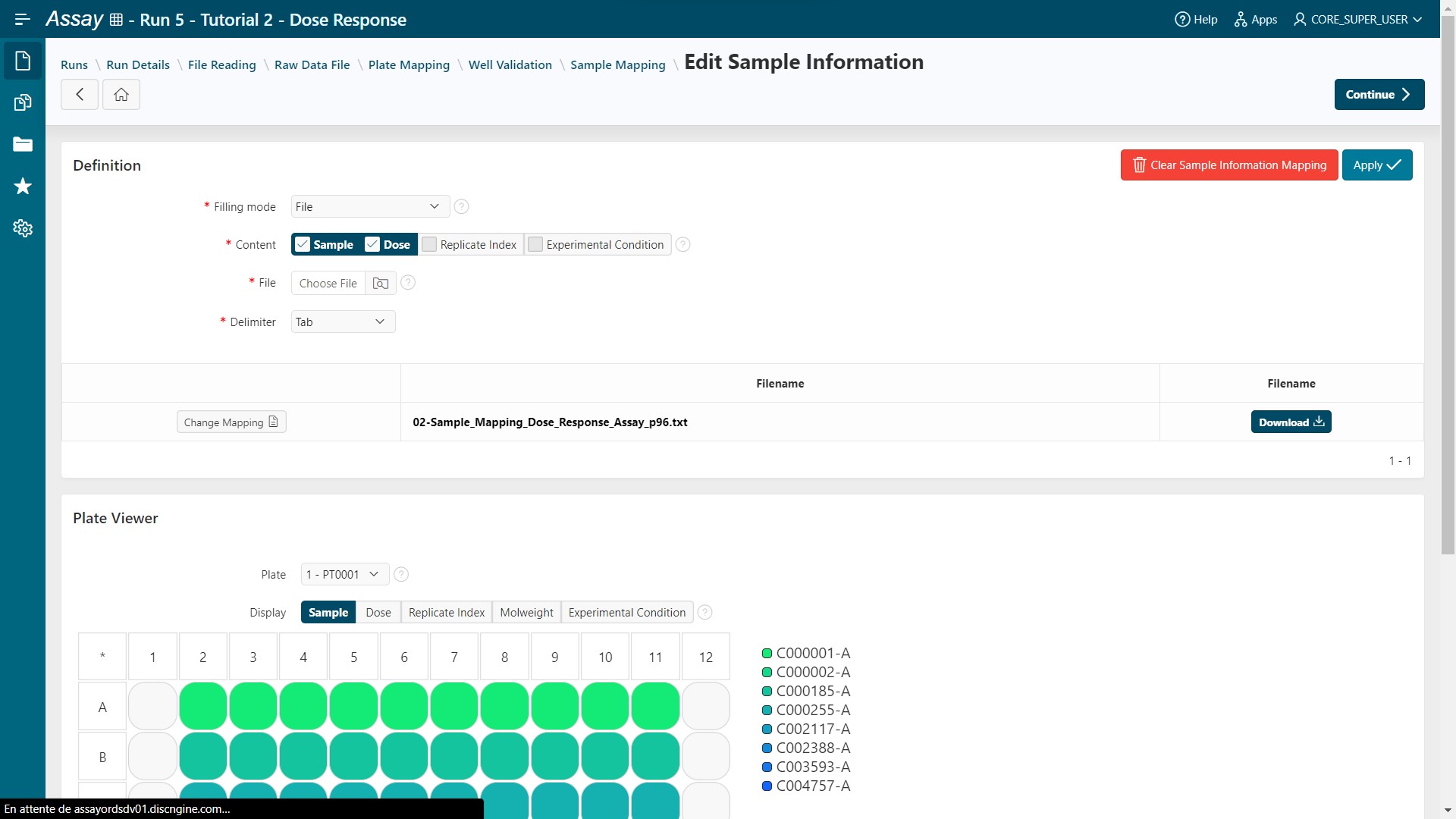
- Click on
Continuebutton.
Samples and doses are mapped for all plates.
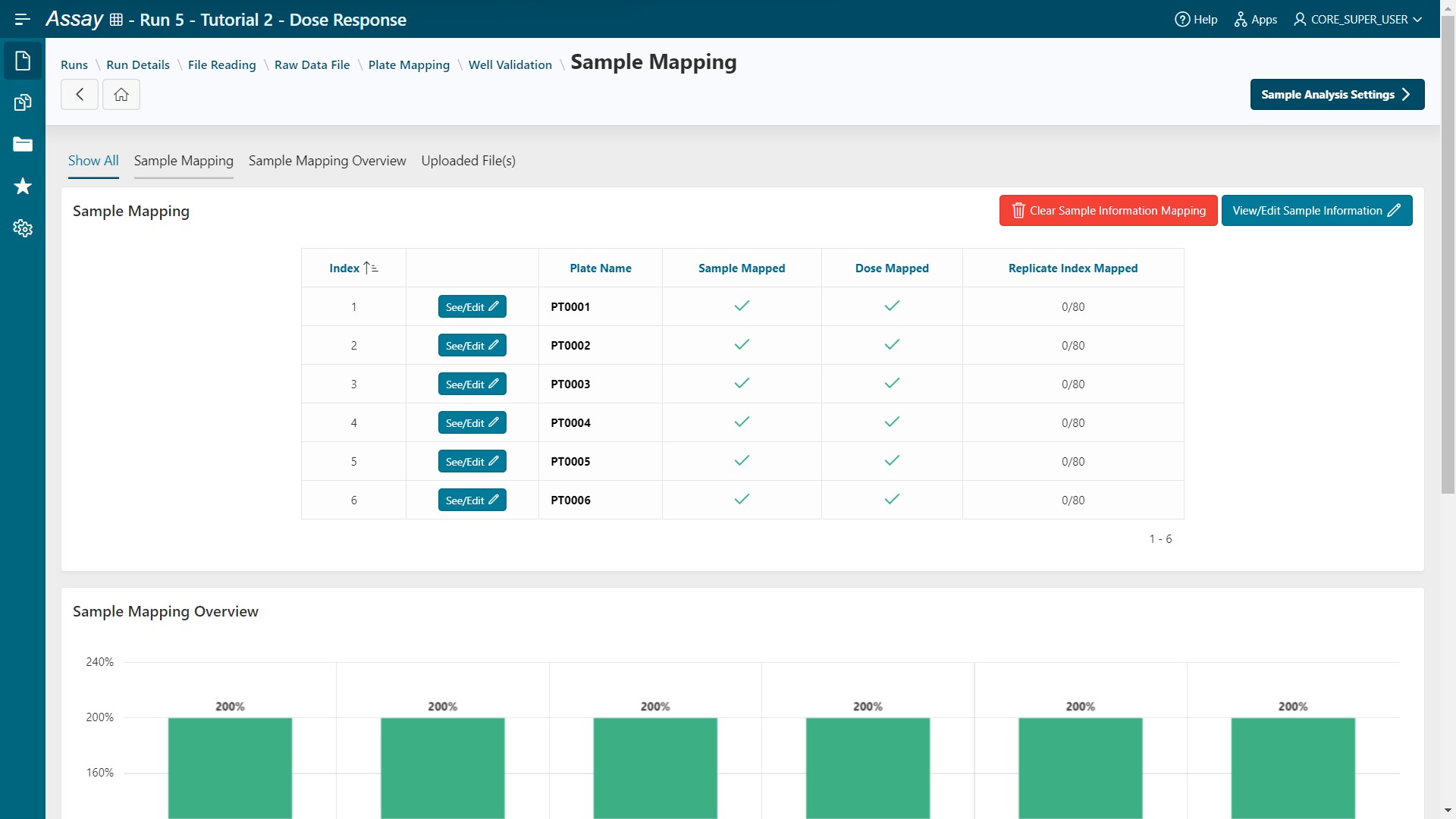
- Click on
Sample Analysis Settingsbutton.
Sample Analysis Settings¶
- Select the parameter to aggregate:
SampleName. - Select the Series:
PCT_INH_READOUT1. - Click on
Create Analysis Groupsbutton: 48 groups are created.
- Select the analysis
Dose Response Fitting, then click on theAdd Sample Analysisbutton. - Leave the parameters empty, click on
Saveto apply the method to all series.
- Go to the Sample Analysis step.
Sample Analysis¶
Select your Visualization Tool:
- TIBCO Spotfire Webplayer
- Pipeline Pilot
Info
This step is only available if 2 visualization tools are installed. Otherwise, this step will be skipped.
- Apply an existing template
- Dose Response Fitting (Spotfire Template):
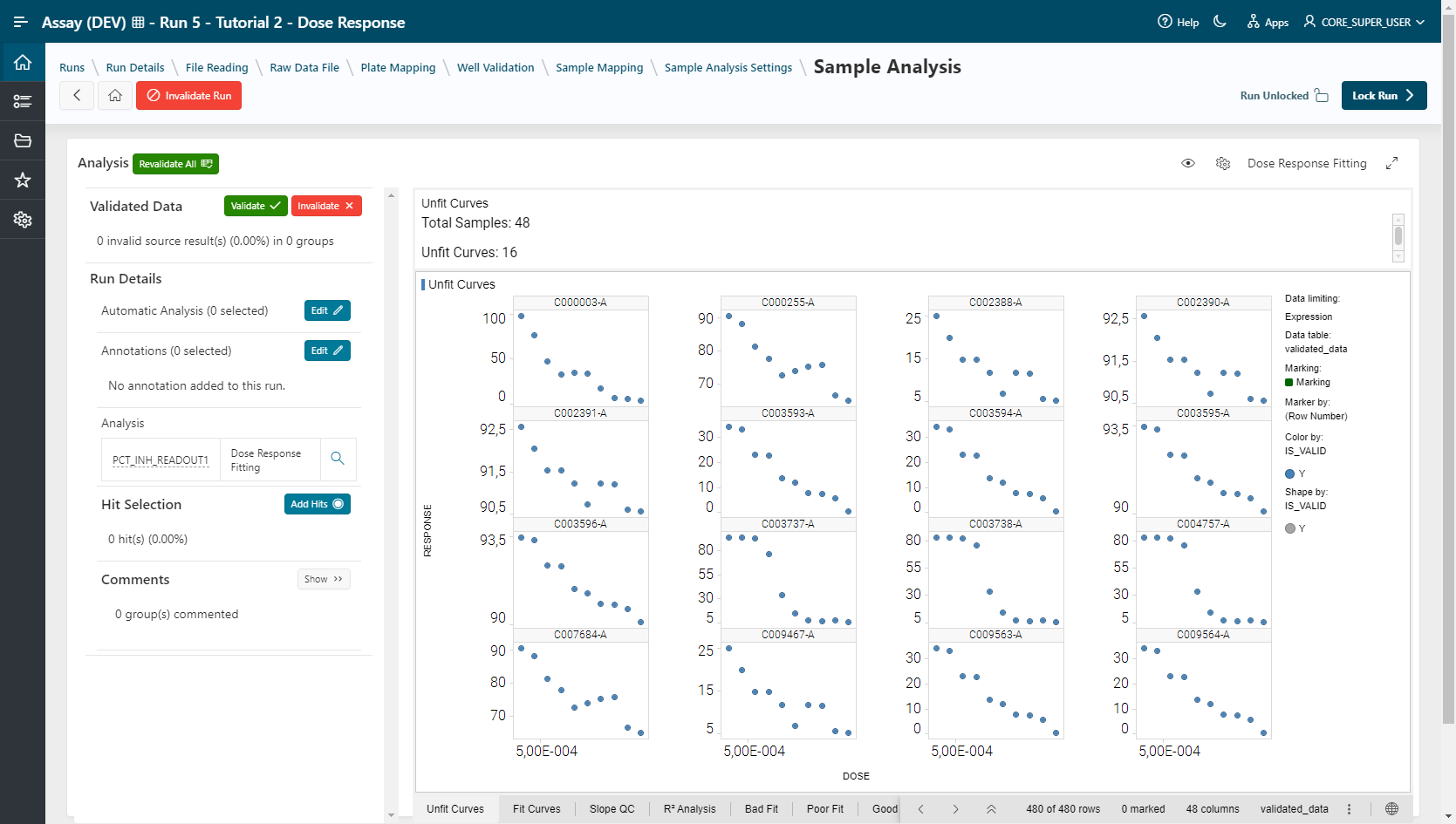
For Hit Selection, select points (hits) in the visualization and add them to a list by clicking on the Add Hits button. This selection can be exported in CSV format (export button) or used to create a list that can be used in the Sample application.
- Lock your Analysis using the
Lock Runbutton. - Go to Publisher step.
Publisher Selection & Workflow Template Creation¶
- Select a target publisher (depending on what is installed – None, Warehouse or BIOVIA Project Data). For more information on publishing data see the Publisher guide.
- Save your analysis by clicking on
Save As Run Template. The template will be used later in the execution mode.
- Give a name and description to the run template.
- Click on
Apply Changesto save your modifications.
Info
Once the run template is saved, the screeners can use it in Execution Mode, following this tutorial